 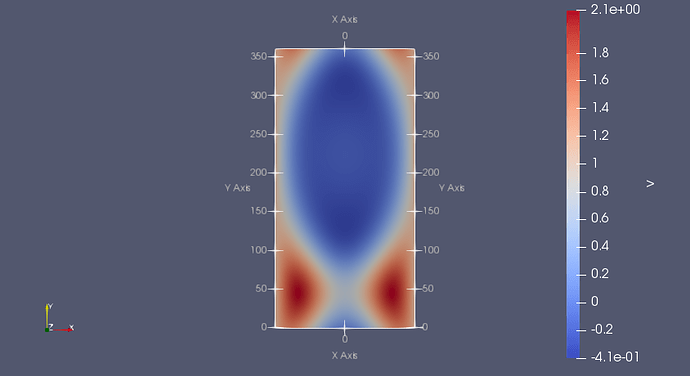
Furthermore, our current application of the reader is the visualization of higher order simulation output which demands for a special representation of the data within a cell. We aim at parallelism to be able to process large data sets. This approach allowed us to tap into the powerful reading and rendering capabilities of ParaView, while the reader is easy to install. The implementation of the UGRID Reader has been designed corresponding to the ParaView plugin architecture. As the UGRID Conventions are increasingly popular with an important subset of the CF community, they warrant the development of a customized tool for the visualization and exploration of UGRID-conforming data. The UGRID Conventions proposed by the UGRID Interoperability group are attempting to fill in this void by extending the CF Conventions with topology specifications. is it a one dimensional network, a 2D triangular mesh or a flexible mixed triangle/quadrilateral mesh, a 2D mesh with vertical layers, or a fully unstructured 3D mesh. 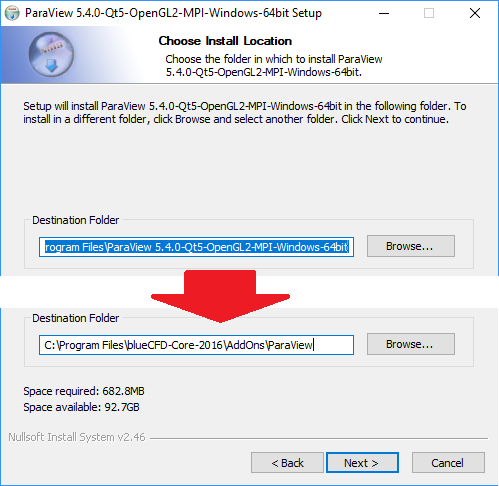
However, it is often necessary to have additional mesh topology information, i.e. While they allow storing unstructured data simply as data defined at a series of points, they do not currently address the topology of the underlying unstructured mesh. The Climate and Forecast Metadata Conventions (CF Conventions) have been set for many years as the standard framework for climate data written in NetCDF format. For this tutorial, I will be using the ParaView example data in Examples netCDF. It currently supports the reading and visualization of 2D unstructured triangular, quadrilateral and mixed triangle/quadrilateral meshes, while the data can be defined per cell or per vertex. Click the open button and find your data. already reached out to you and offered help.We present the UGRID Reader, a visualization software component that implements the UGRID Conventions into Paraview. That reader is maintained by DKRZ and you would have to go through them for help with this reader. That said, on the previous post I thought we determined that the appropriate reader was the CDI reader (available through a plugin). We usually need to have an example file or at a minimum the output of ncdump -h. So simply sending a screenshot is usually not enough information to determine what is going on. nc extension and expect consumers to magically understand what they mean.īecause there is this ambiguity on the format of data in netCDF files, there are many different netCDF readers to handle the different known conventions. It kind of drives me nuts that so many netCDF producers don’t recognize this fact and just write out files with a. Thankfully, different ASCII file producers with different conventions tend to understand are each a format in its own right and identify the file as such. 
csv) or a hierarchy with tags describing polygonal data (vtp). 
For example, an ASCII file could be a table with delimiters (i.e. You need to specify a convention of how the data are arranged. That’s like saying “I have an ASCII file.” That could mean lots of different things. So simply saying “I have a netCDF file” is not sufficient information for ParaView to read it. That alone is not enough for a ParaView reader to just know how to read the data. NetCDF really just holds a collection of multi-dimensional arrays with attributes like a name. To back up a step, it’s important to note that the netCDF file format is not really a file format from the perspective of ParaView or other data consumers. Didn’t we just hash this out on this post? 
0 Comments
Leave a Reply. |
Details
AuthorWrite something about yourself. No need to be fancy, just an overview. ArchivesCategories |
 RSS Feed
RSS Feed
